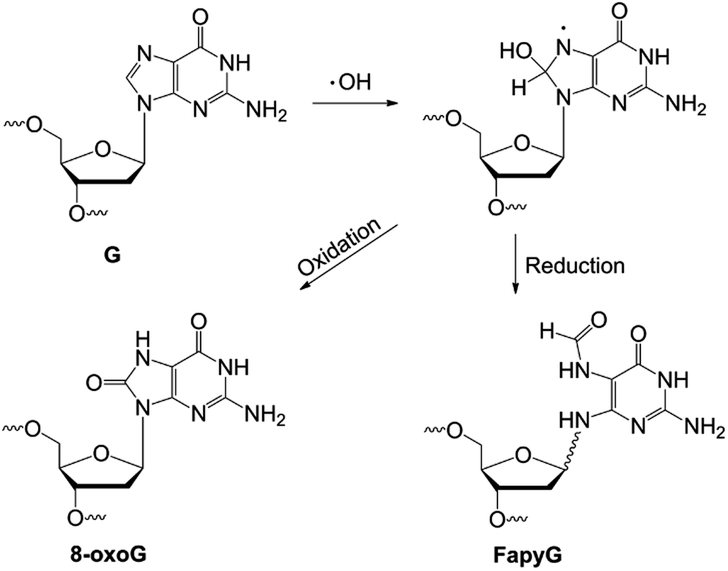
Molecular beacons with oxidized bases report on substrate specificity of DNA oxoguanine glycosylases - Chemical Science (RSC Publishing) DOI:10.1039/D1SC05648D

Rational design of highly potent broad-spectrum enterovirus inhibitors targeting the nonstructural protein 2C. - Abstract - Europe PMC

Exploiting genomic surveillance to map the spatio-temporal dispersal of SARS-CoV-2 spike mutations in Belgium across 2020 | Scientific Reports
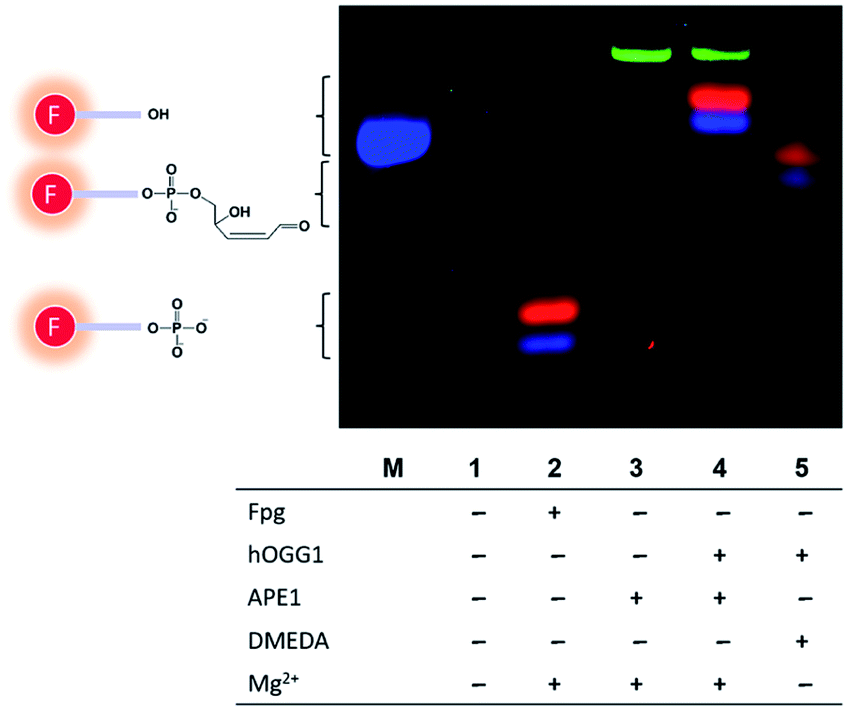
Molecular beacons with oxidized bases report on substrate specificity of DNA oxoguanine glycosylases - Chemical Science (RSC Publishing) DOI:10.1039/D1SC05648D

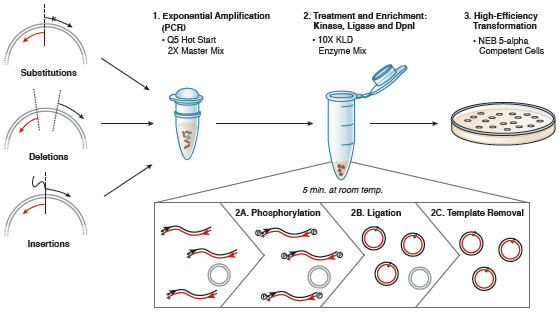
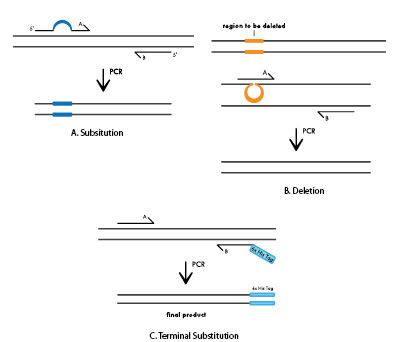
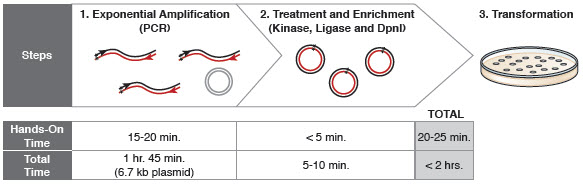
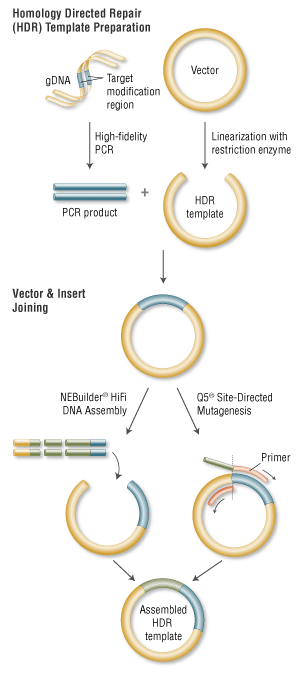
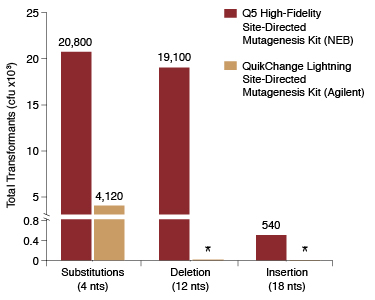
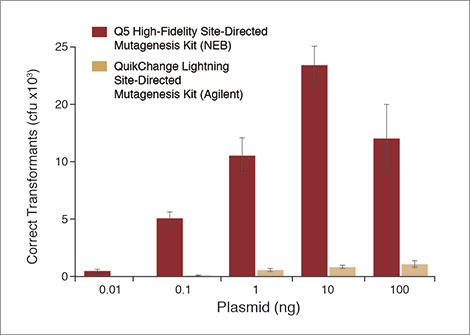
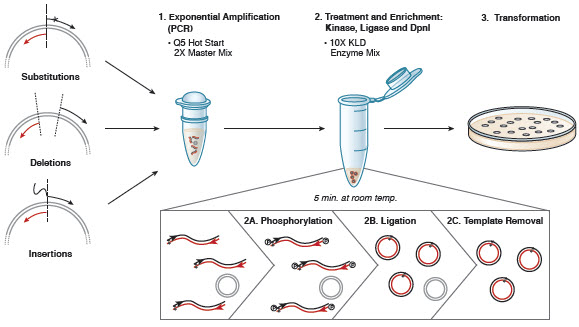
Q5 ® Site-Directed Mutagenesis Kit. Reprinted from www.neb.com (2013)... | Download Scientific Diagram
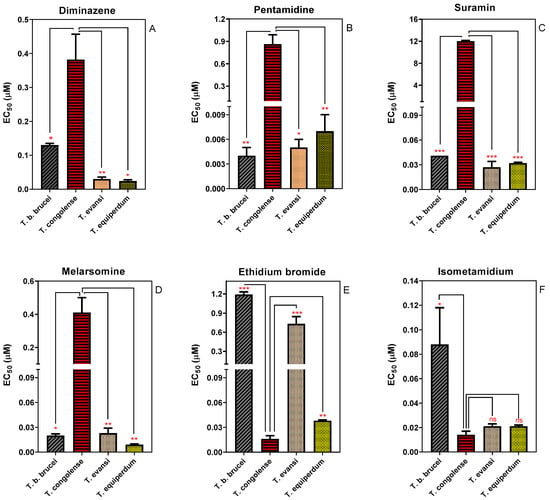
IJMS | Free Full-Text | Differences in Transporters Rather than Drug Targets Are the Principal Determinants of the Different Innate Sensitivities of Trypanosoma congolense and Trypanozoon Subgenus Trypanosomes to Diamidines and Melaminophenyl
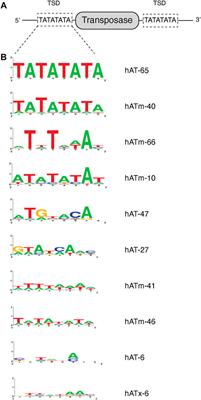
Frontiers | Insertion Specificity of the hATx-6 Transposase of Hydra magnipapillata | Molecular Biosciences

A Proteomic Study on the Membrane Protein Fraction of T Cells Confirms High Substrate Selectivity for the ER Translocation Inhibitor Cyclotriazadisulfonamide - Molecular & Cellular Proteomics